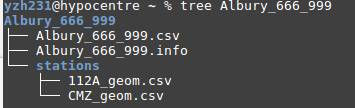
Please read the readme @ https://github.com/ucgmsim/IM_calculation/blob/master/README.md for instructions on how to run the code.
Tested on very simple multi-process on Kupe with good speed-up using 40 and 80 cores. For the sample, 2228 stations were used
| Machine | Cores | Time |
|---|---|---|
| Hypocentre | 1 | 132m |
| Hypocentre | 8 | 8.7m |
| Kupe | 40 | 27m |
| Kupe | 80 |
With command : python calculate_ims.py ../BB.bin b -o /home/yzh231/ -i Albury_666_999 -r Albury -t s -v 18p3 -n 112A CMZ -m PGV pSA -p 0.02 0.03 -e -c geom -np 2
The result is outputted to the following location, where:

All the steps below are to be carried out in hypocentre
The following steps should only be performed once for each selected binary file
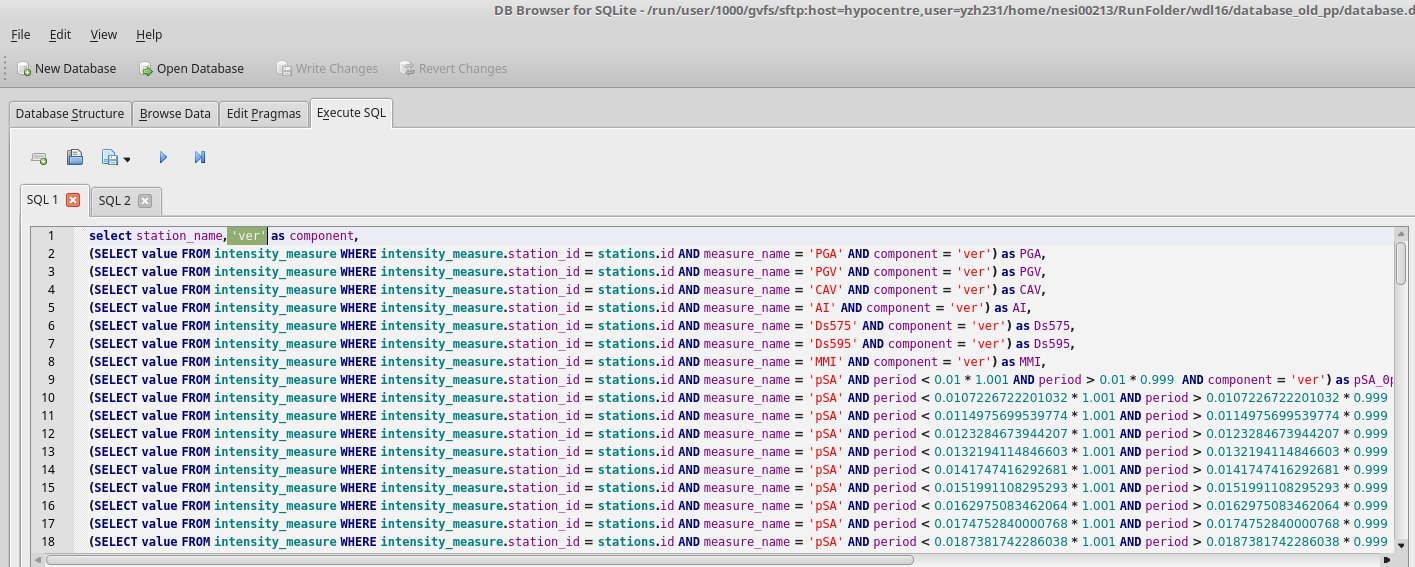
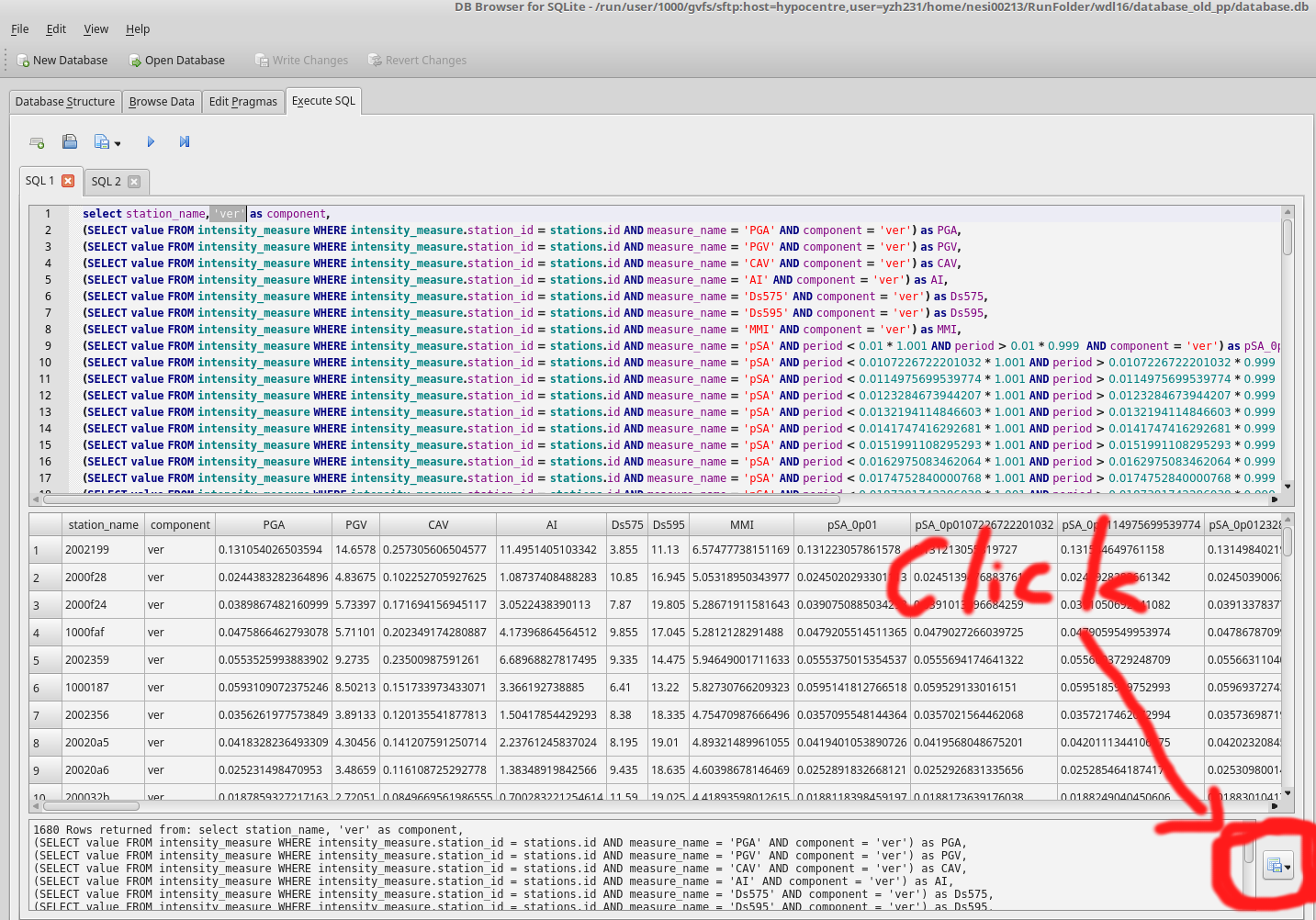

2.Generate test input files
Follow the instruction in Binary Workflow FAQ, we can generate single waveform files. These waveforms are intended for the testing of ascii functionality of calculate_ims.py. Open a python cell
from qcore.timeseries import BBSeis
bb = BBSeis('/nesi/transit/nesi00213/RunFolder/daniel.lagrava/Kelly_VMSI_Kelly-h0p4_EMODv3p0p4_180531/BB/Cant1D_v2-midQ_leer_hfnp2mm+_rvf0p8_sd50_k0p045/Kelly_HYP01-03_S1244/Acc/BB_with_siteamp.bin')
bb.all2txt(self, prefix='/home/$user/benchmark_im_sim_waveforms/', f='acc'): |
Now we have all the wavefoms. 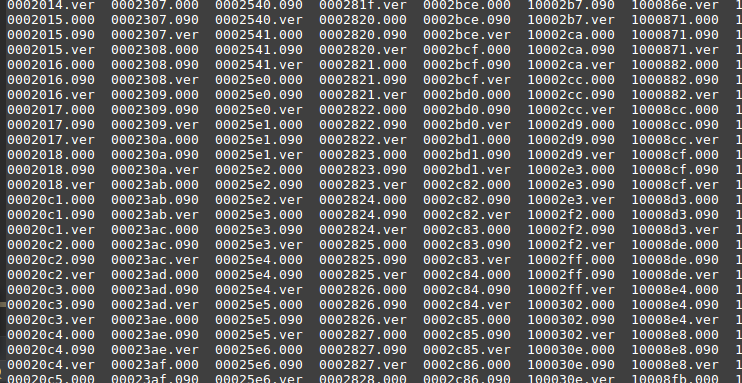
NOW you have all the input files ready
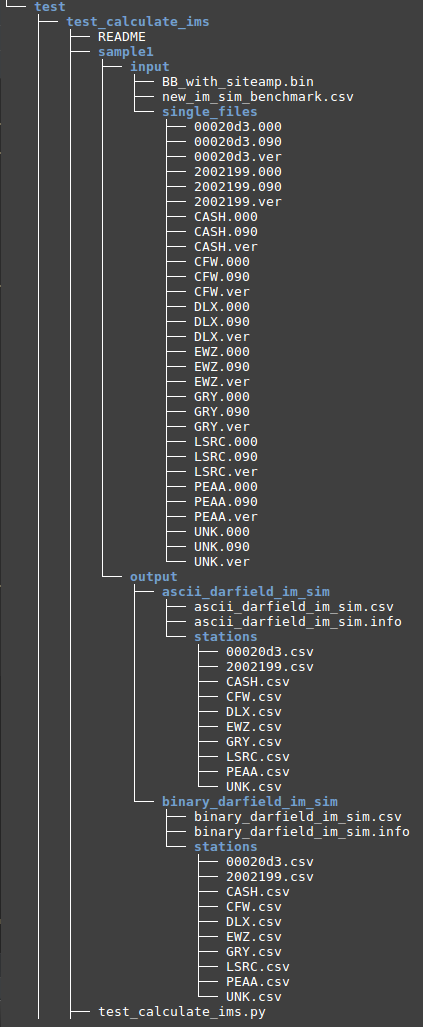
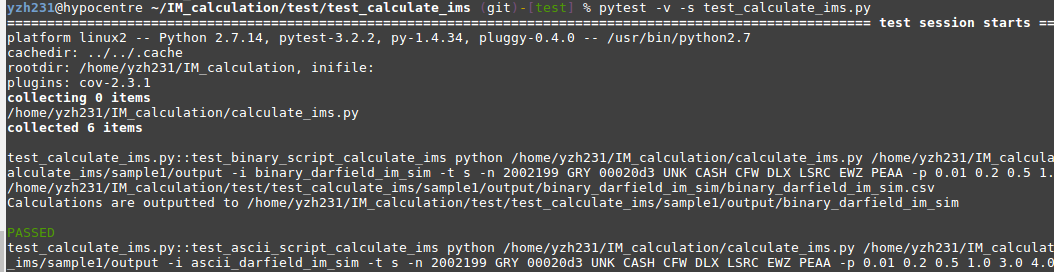
Make sure you are currently under the test_calculate_ims folder, run: pytest -v -s test_calculate_ims.py
Checkpointing is needed for IM_calculation due to large job size and limited running time on Kupe. Therefore, we implemented checkpointing to track the current progress of an im_calculation job, and carry on from where the job was interrupted by slurm.
To run checkpointing, first git clone the IM_calculaiton repo, then check out the checkpoint branch
$ git clone https://github.com/ucgmsim/IM_calculation.git $ git checkout checkpoint |
Open im_calc_sl.template, change the IMPATH variable (line 22) to where you have cloned the git repository
# open template ~/IM_calculation-[checkpoint]$ vim im_calc_sl.template # modify the $IMPATH export IMPATH=/home/melody.zhu/IM_calculation |
Note, the checkpointing code relies on the input/output directory structure specified in the im_calc_al.template in the checkpoint branch. Failure to match the dir structure will result in runtime error. A quick fix would be modifying the template to suit your own dir structure.
(1) Simulation
Input/output structure defined in im_calc_al.template

Actual input data structure:

The input binary file is under:
/nesi/nobackup/nesi00213/RunFolder/Cybershake/v18p6_batched/v18p6_1k_under2p0G_ab/Runs/BlueMtn/BB/Cant1D_v3-midQ_OneRay_hfnp2mm+_rvf0p8_sd50_k0p045/BlueMtn_HYP28-31_S1514/Acc/BB.bin |
The output IM_calc folder is under:

(2) Observed
Input/output structure defined in im_calc_al.template

Actual input data structure:

The output IM_calc folder is under:

Sample command to run checkpointing:(use generate_sl2.py)
$ python generate_sl2.py -s /nesi/nobackup/nesi00213/RunFolder/Cybershake/v18p6_batched/v18p6_1k_under2p0G_ab/Runs -srf /nesi/nobackup/nesi00213/RunFolder/Cybershake/v18p6/Runs/test_srfs/ -ll /scale_akl_nobackup/filesets/transit/nesi00213/StationInfo/non_uniform_whole_nz_with_real_stations-hh400_v18p6.ll -np 80 -o ~/test_obs/IMCalcExample /nesi/nobackup/nesi00213/RunFolder/Cybershake/v18p6/test_check_point/ > test.sl |
To submit the slurm script:
$cp test.sl /nesi/nobackup/nesi00213/tmp/auto_preproc $sbatch test.sl |
The reason that we have to run test.sl under '/nesi/nobackup/nesi00213/tmp/auto_preproc' is otherwise slurm cannot find machine.env specified by the test.sl script:
Splitting a big slurm script into several smaller slurms is need due to maximum number of lines allowed in a slurm script on Kupe.
Still in the checkpointing branch, Run the generate_sl3.py script that uses both checkpointing and splitting. The -ml argument specifies the maximum number of lines of python call to calculate_ims.py/caculate_rrups.py. Header and footer like #SBATCH --time=15:30:00, date etc are not included.
Say if the max number of lines allowed in a slurm script is 1000, and your (header + footer) are 30 lines, then the number n that you pass to -ml should be 0 < n <=967. eg. -ml 967.
We have 250 simulation dirs to run, by specifying -ml 100 (100 python calls to calculate_ims.py per slurm script), we expect 3 sim slurm scripts to be outputted.(1-100, 100-200, 200-250)
We have 3 observed dirs to run, by specifying -ml 100 (100 python calls to calculate_ims.py per slurm script), we expect 1 sim slurm scripts to be outputted.
We have 61 rrup files to run, by specifying -ml 100 (100 python calls to calcualte_rrups.py per slurm script), we expect 1 sim slurm scripts to be outputted.
Command:
python generate_sl3.py -s /nesi/nobackup/nesi00213/RunFolder/Cybershake/v18p6_batched/v18p6_1k_under2p0G_ab/Runs -srf /nesi/nobackup/nesi00213/RunFolder/Cybershake/v18p6/Runs/test_srfs/ -ll /scale_akl_nobackup/filesets/transit/nesi00213/StationInfo/non_uniform_whole_nz_with_real_stations-hh400_v18p6.ll -np 80 -o ~/test_obs/IMCalcExample/ /nesi/nobackup/nesi00213/RunFolder/Cybershake/v18p6/test_check_point/ -ml 100 |
Output: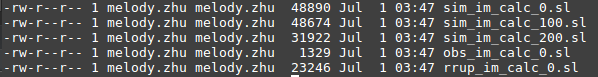
Todo: tidy up check_point.py, generate_sl[*|2|3].py